Overview
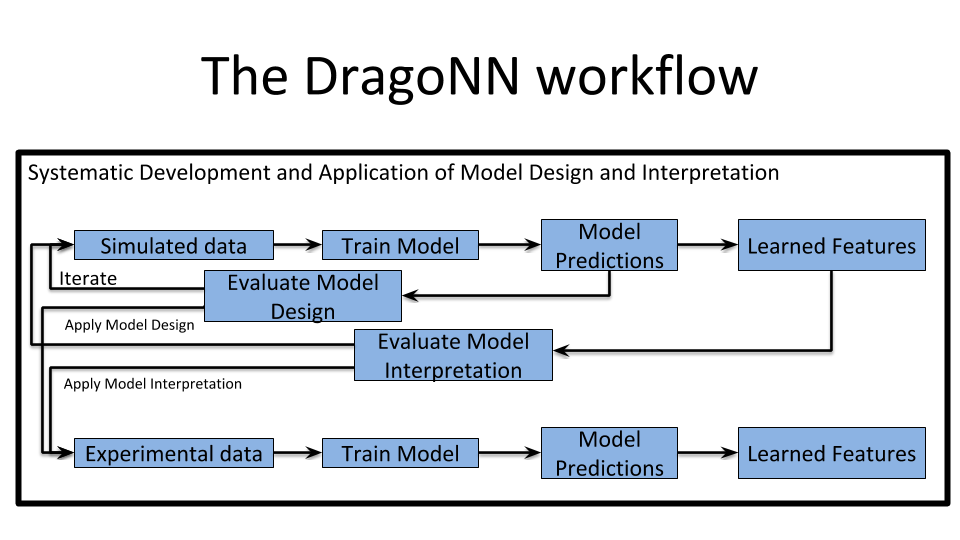
Web-based DragoNN tutorials use Jupyter notebooks to allow interactive model manipulation and visualization. The DragoNN tutorials demonstrate model design and interpretation principles using simulated data. Having access to the ground truth in the simulations enables us to dissect the specific parameters affecting prediction performance and model interpretation. Simulations can thus enable systematic evaluation of model design and interpretation to quickly find principles that could effectively apply to real genomic data.
Getting access to DragoNN software and GPU hardware
Each tutorial below specifies software and hardware requirements.
Tutorials with software requirements will necessitate either a released version of the DragoNN package or the bleeding-edge (non-released) version. See the code page for software installation and configuration instructions.
Tutorials with hardware requirements will necessitate either a CPU (Tutorial 1) or a GPU (Tutorials 2 - 5). If the tutorial requires a GPU and you do not have access to GPUs locally, you can run the tutorials in Google Colaboratory (see below) with a GPU accelerator.
Google Colaboratory Tutorials
The following notebooks can be opened and executed with Google Colaboratory
Tutorial 2: CNN hyperparameter tuning via grid search.
Tutorial 3: Interpreting features induced by CNN's across multiple types of motif grammars.
Tutorial 5: Functional variant characterization for non-coding SNPs within the SPI1 motif.
Intro tutorials included with the dragonn source code
Software requirements: bleeding-edge DragoNN package.
Hardware requirements: CPU/laptop sufficient for tutorial 1. GPU hardware required for tutorials 2 - 5.
To explore the Jupyter notebook tutorials, navigate to the ./tutorials directory and run
jupyter notebook
This will start a jupter notebook server, allowing you to navigate to the tutorial notebooks in your browser:
- Exploring convolutional neural network (CNN) architectures for simulated genomic data
- CNN Hyperparameter Tuning via Grid Search
- Interpreting features induced by DNN's across multiple types of motif grammars
- Interpreting predictive sequence features in TF binding events within the GM12878 cell line.
- Functional variant characterization for non-coding SNPs within the SPI1 motif